 In other words, in half of the molecules H a is shielded by H b (thus the NMR signal is shifted slightly upfield) and in the other half H a is deshielded by H b(and the NMR signal shifted slightly downfield).\). This shift in the NMR frequency due to the electronic molecular orbital coupling to the external magnetic field is called chemical shift, and it explains. The B eff ‘felt’ by H a is a slightly weaker if H b is aligned against B 0, or slightly stronger if H b is aligned with B 0. A standard iNMR window, enriched with chemical shift labels under the spectrum A drawer at its side, containing the parameters specific to simulation A dialog (sheet) where the user specifies the number and kind of systems, nuclei and parameters. The magnetic moment of H b will be aligned with B 0 in (slightly more than) half of the molecules in the sample, while in the remaining half of the molecules it will be opposed to B 0. Here's how it works, looking first at the H a signal: in addition to being shielded by nearby valence electrons, each of the H a protons is also influenced by the small magnetic field generated by H b next door (remember, each spinning proton is like a tiny magnet). Lecture 3: Coupling Scenario 2:protonAis adjacent (vicinal)to two protons, and C, but B and C havethe same chemical Now, we get a triplet: Thedouble intensity forthemiddle line comes there are two permutations. The DIET (or dual interval echo train) sequence, a modification of the fast spin echo (FSE) sequence that selectively reduces signal from fat in MR images. Here, we directly look into the 15 N-15 N homonuclear J-coupling network in the system and use uniformly 13 C and 15 N labeled histidine samples at pH 6.3 and 11.0 as an example to examine whether or not there exists a two-bond J-coupling between N 1 and N 2 (i.e. In our 1,1,2 trichloromethane example, the H a and H b protons are spin-coupled to each other. Carbon-13 chemical shifts and the POC, POCC, PNC and PNCC coupling constants of 18 compounds containing the amine moiety, and with the general formula Y2P (X)NHR YC2H5O, C6H5O, CH2O. The C C coupling constants in the aromatic molecule benzene as well as C H. Here, we apply an orbital analysis to one-bond carboncarbon coupling, J(C,C), in ethane, ethene, and ethyne.  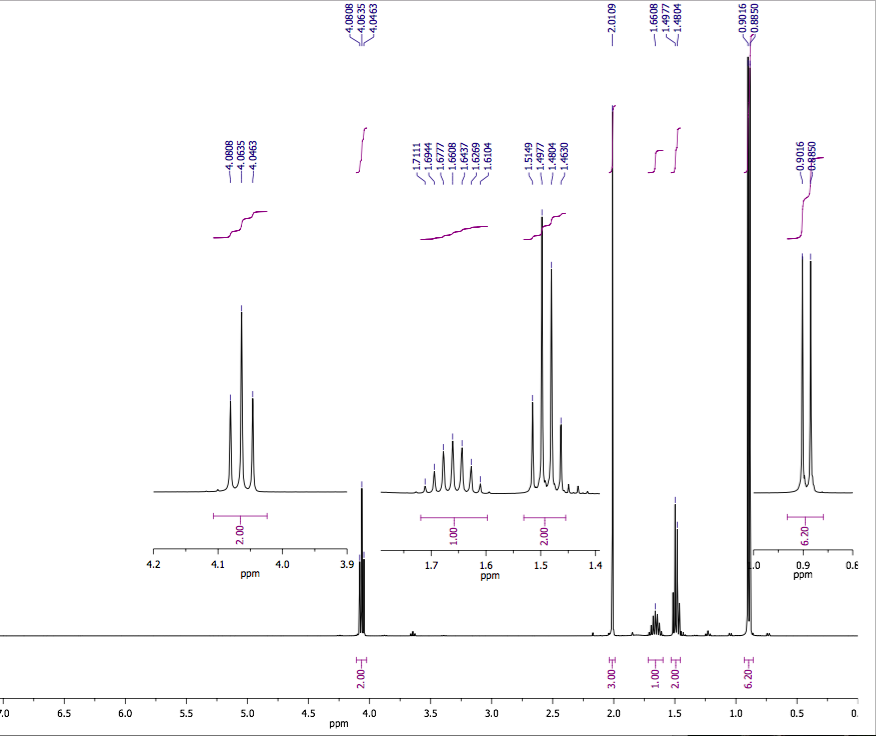 The source of signal splitting is a phenomenon called spin-spin coupling, a term that describes the magnetic interactions between neighboring, non-equivalent NMR-active nuclei. We should point out that such analyses for organic (1217) and inorganic systems (1822) are also an active research topic. Smaller shifts of either sign can occur further out. When its time to specify the first J he is soon in trouble. The Calculation of Indirect Nuclear SpinSpin Coupling Constants 102. One may therefore be tempted to put them into the same row. The H b signal at 5.76 ppm, on the other hand, is split into three sub-peaks, with the middle peak higher than the two outside peaks - if we were to integrate each subpeak, we would see that the area under the middle peak is twice that of each of the outside peaks. Nuclei A and D have identical chemical shifts and identical coupling constants. The signal at 3.96 ppm, corresponding to the two H a protons, is split into two subpeaks of equal height (and area) – this is referred to as a doublet. outfile File to write j-coupling values to with fixed format. jcoupling outfile kfile out name Atom mask in which to search for dihedrals within.iNMR (evolution of SwaN-MR) : An NMR processing software for the Macintosh. In this and in many spectra to follow, we show enlargements of individual signals so that the signal splitting patterns are recognizable. jcoupling Calculate J-coupling values from specified dihedral angles. ANSIG Assignment of NMR Spectra by Interactive Graphics (written in Fortran. Rather than being a complication, however, this splitting behavior actually provides us with more information about our sample molecule.Ĭonsider the spectrum for 1,1,2-trichloroethane. of Residual Dipolar Couplings in MR Spectra and Data Analysis of Carnosine. 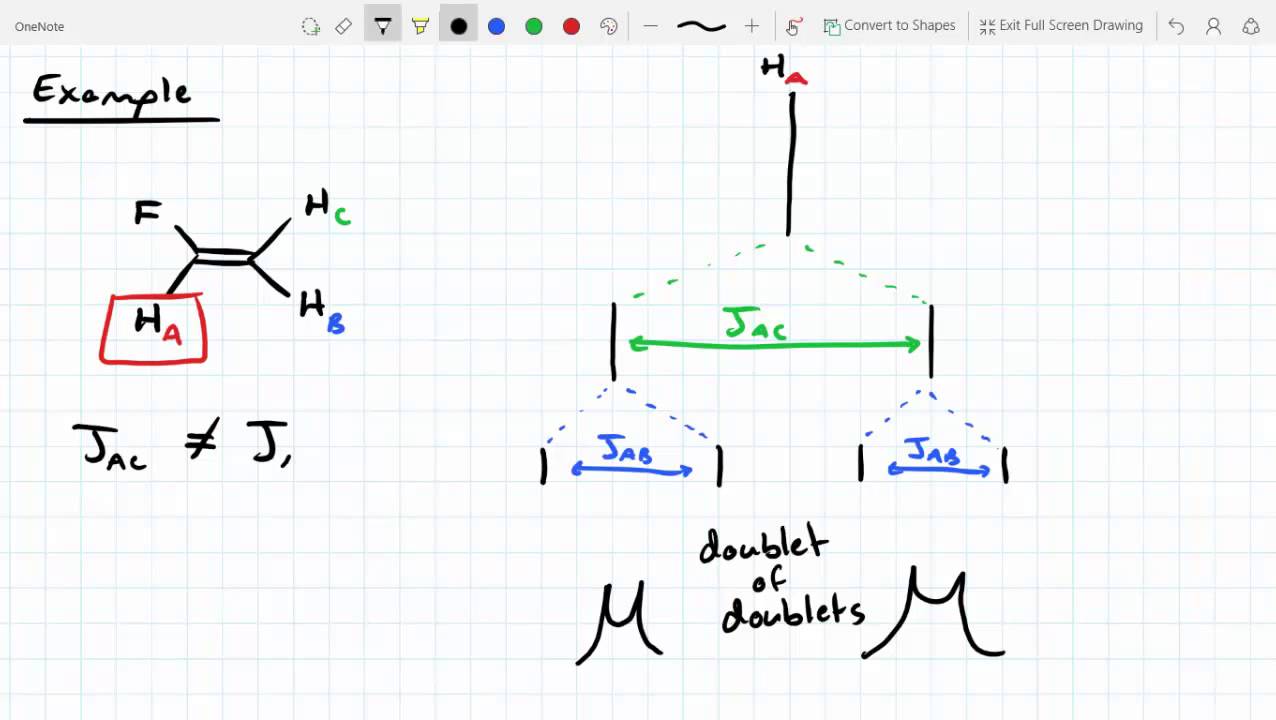 In fact, the 1H-NMR spectra of most organic molecules contain proton signals that are 'split' into two or more sub-peaks. Peak splitting due to the residual dipolar coupling (RDC) represents a. The 1H-NMR spectra that we have seen so far (of methyl acetate and para-xylene) are somewhat unusual in the sense that in both of these molecules, each set of protons generates a single NMR signal.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
